Using ettbc: Emulating a Target Trial for Breast Cancer Screening
2026-05-04
1 Background
This article demonstrates how to use the {ettbc} package to emulate a target trial for breast cancer screening using the clone-censor-reweight methodology. The methods are based on the study by García-Albéniz et al. (2020), who estimated the effect of continuing annual screening mammography on breast cancer mortality in Medicare beneficiaries aged 70–84 years.
The Target Trial
The study aimed to answer: Does continuing annual screening mammography (compared with stopping) reduce breast cancer mortality in older women?
Because a randomised trial of this question is unlikely to be conducted, the target trial was emulated using Medicare claims data. The key methodological challenge is confounding by indication: women who continue screening tend to be healthier and more health-conscious than those who stop.
The Clone-Censor-Reweight Approach
The clone-censor-reweight method handles time-varying treatment and censoring in three steps (Hernán and Robins 2016):
Clone: Each participant is replicated into two copies — one assigned to the STOPBASE arm (stop screening at study entry) and one to the CONTINUE arm (continue annual screening).
Censor: Clones are censored in their assigned arm when they deviate from the protocol:
- STOPBASE: censored at the first screening mammogram received after the initial grace period.
- CONTINUE: censored when more than 14 months pass without any mammogram.
Reweight: Inverse probability weights (IPW) are constructed to account for the informative censoring introduced in step 2.
The {ettbc} package implements the first two steps with clone_censor() and provides a helper function expand_to_long() to prepare the dataset for the IPW outcome analysis.
2 Example Data
The package ships with three synthetic datasets that mimic the structure of the original study data (but contain entirely simulated values).
Months are numbered consecutively from 1 (January 2000) to 108 (December 2008). Participants enter the study between months 1 and 60.
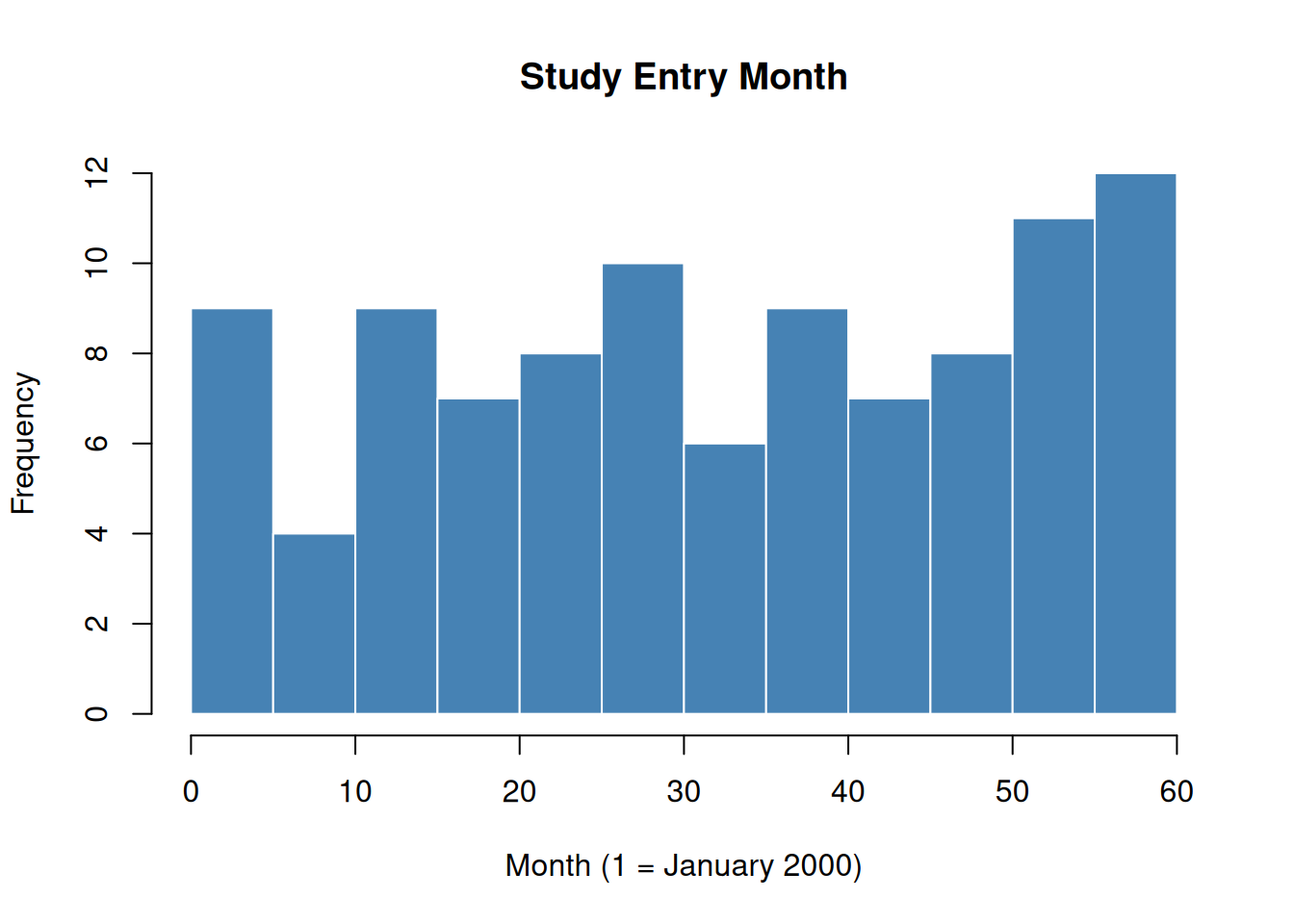
Figure 1: Distribution of study entry months in the simulated cohort.
3 Step 1: Clone and Censor
clone_censor() takes the cohort and mammography event datasets and returns a data frame with two rows per participant — one for each arm.
Code
cloned <- clone_censor(
cohort,
screening_mammograms,
diagnostic_mammograms
)
nrow(cloned) # should be 2 × nrow(cohort)
#> [1] 200
head(cloned[, c("id", "arm", "start_month", "end_month",
"censor_month", "fup", "died", "bc_died")])
#> id arm start_month end_month censor_month fup died bc_died
#> 1 1 STOPBASE 26 38 38 13 0 0
#> 2 2 STOPBASE 33 45 45 13 0 0
#> 3 3 STOPBASE 28 42 42 15 0 0
#> 4 4 STOPBASE 1 12 12 12 0 0
#> 5 5 STOPBASE 17 28 28 12 0 0
#> 6 6 STOPBASE 40 52 52 13 0 0The key new columns are:
arm: trial arm ("STOPBASE"or"CONTINUE")end_month: final observed month (minimum of death, administrative censoring, and protocol censoring)censor_month: month of protocol deviation censoring (NAif not censored for protocol non-adherence)fup: follow-up time in monthsdied: overall mortality indicator (1 if death caused end of follow-up)bc_died: breast cancer mortality indicator
Censoring Patterns by Arm
Because participants in STOPBASE are censored when they get a screening mammogram, annual screeners are censored early in that arm. Conversely, participants in CONTINUE are censored when they go too long without a mammogram, so non-adherers are censored early.
Code
par(mfrow = c(1, 2))
stopbase <- cloned[cloned$arm == "STOPBASE", ]
continue <- cloned[cloned$arm == "CONTINUE", ]
hist(stopbase$fup,
breaks = 20, main = "STOPBASE",
xlab = "Follow-up (months)", col = "#E05C4B", border = "white",
xlim = c(0, 110)
)
hist(continue$fup,
breaks = 20, main = "CONTINUE",
xlab = "Follow-up (months)", col = "#4B8FE0", border = "white",
xlim = c(0, 110)
)
par(mfrow = c(1, 1))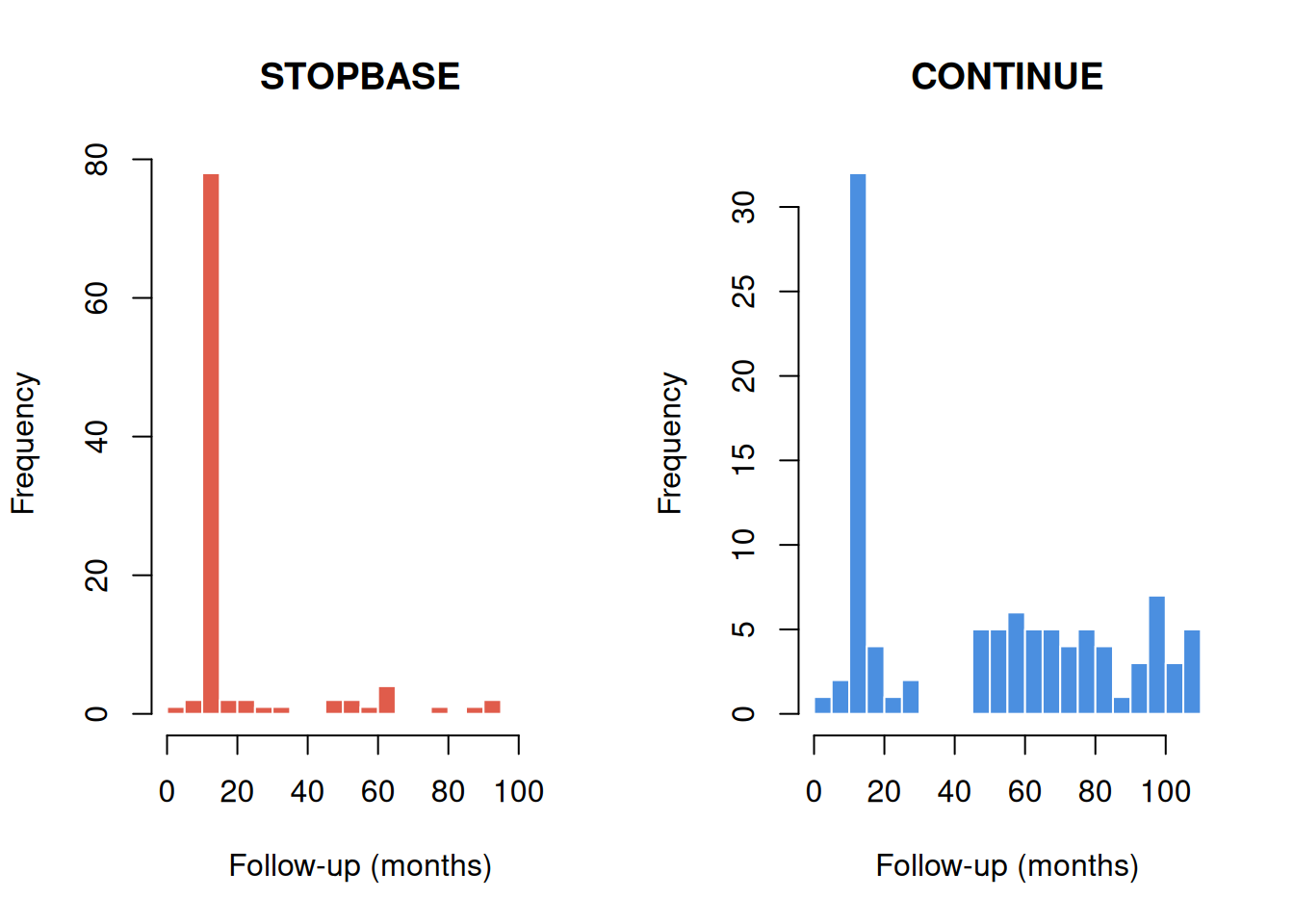
Figure 2: Follow-up time by arm and censoring type.
As expected, most STOPBASE clones are censored (those who continued getting annual mammograms), while only a minority of CONTINUE clones are censored (those who stopped getting mammograms).
4 Step 2: Expand to Long Format
expand_to_long() converts the cloned dataset to one row per participant-arm-month. This format is required for the discrete-time survival models used to estimate the outcome.
Code
long_data <- expand_to_long(cloned)
nrow(long_data) # total person-arm-months
#> [1] 7052
head(long_data)
#> id arm month month2 dead_t1 bc_dead_t1 bc_long
#> 1 1 STOPBASE 26 0 0 0 0
#> 2 1 STOPBASE 27 1 0 0 0
#> 3 1 STOPBASE 28 2 0 0 0
#> 4 1 STOPBASE 29 3 0 0 0
#> 5 1 STOPBASE 30 4 0 0 0
#> 6 1 STOPBASE 31 5 0 0 0The key columns in the long dataset are:
month: calendar month (1–108)month2: months since study entry (0-indexed)dead_t1: death in the next interval:1= yes,0= no,NA= censoredbc_dead_t1: breast cancer death in the next intervalbc_long: breast cancer diagnosis flag at this month
Verifying the Long Format
A simple check: the number of observed deaths in the long dataset should equal the number of died == 1 rows in the cloned dataset.
5 Step 3: Descriptive Analysis
Before fitting weighted outcome models, it is useful to describe the data and examine crude survival by arm.
Code
# Compute monthly event rate per arm
arms <- c("STOPBASE", "CONTINUE")
cols <- c("#E05C4B", "#4B8FE0")
plot(NA,
xlim = c(0, 70), ylim = c(0, 0.15),
xlab = "Months since study entry",
ylab = "Cumulative mortality (crude)",
main = "Crude cumulative mortality by arm"
)
for (j in seq_along(arms)) {
sub <- long_data[long_data$arm == arms[j], ]
# Month-by-month event counts
max_t <- max(sub$month2)
cum_mort <- numeric(max_t + 1L)
n_risk <- numeric(max_t + 1L)
for (t in 0:max_t) {
rows_t <- sub[sub$month2 == t, ]
n_risk[t + 1L] <- nrow(rows_t)
cum_mort[t + 1L] <- sum(rows_t$dead_t1 == 1L, na.rm = TRUE)
}
cum_mort <- cumsum(cum_mort) / max(n_risk, 1L)
lines(0:max_t, cum_mort, col = cols[j], lwd = 2)
}
legend("topleft",
legend = arms, col = cols, lwd = 2, bty = "n"
)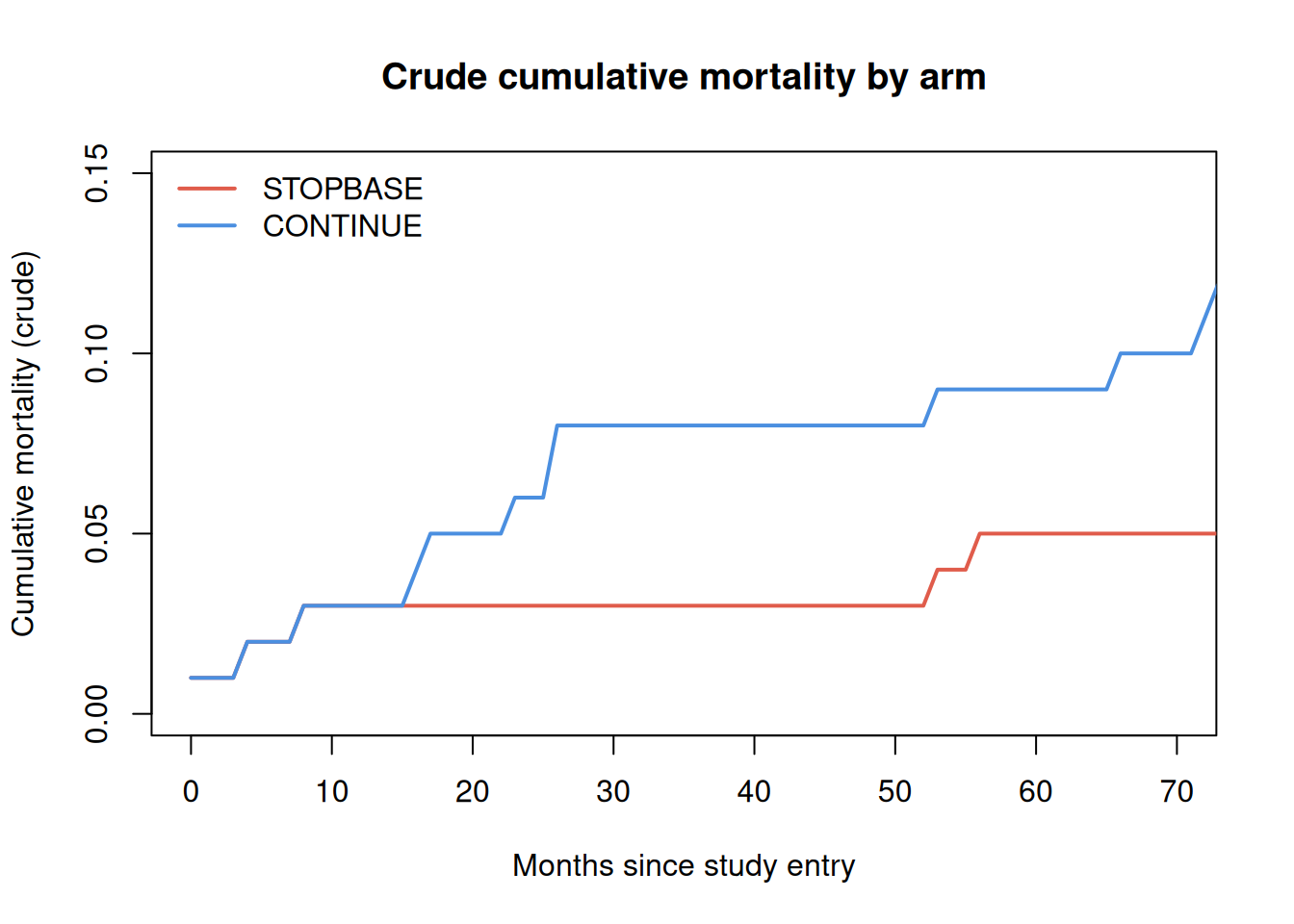
Figure 3: Empirical cumulative mortality by arm (crude, without IPW). Under the null, the two arms should be similar because clones are derived from the same participants.
Note
The crude curves above do not account for the informative censoring introduced by the clone-censor step. In a complete analysis, inverse probability weights (IPW) would be applied to correct for the different censoring mechanisms in each arm. See García-Albéniz et al. (2020) for the full weighted analysis.
6 Summary
The {ettbc} package provides two core functions for the clone-censor step of target trial emulation:
| Function | Input | Output |
|---|---|---|
clone_censor() |
cohort + mammogram events | two rows per participant |
expand_to_long() |
cloned dataset | one row per participant-arm-month |
After running these steps, the long dataset can be used with standard weighted logistic or pooled logistic regression to estimate the intention-to-treat effect of each screening strategy.